Conformalized Forecasting using Machine Learning (and statistical) models with ARCH effects
mlarchf.RdConformalized Forecasting using Machine Learning (and statistical) models with ARCH effects
mlarchf(
y,
h = 10L,
mean_model = forecast::auto.arima,
model_residuals = forecast::meanf,
fit_func = ahead::ridge,
predict_func = predict,
conformal = FALSE,
lags_vol = 10L,
type_pi = c("surrogate", "bootstrap", "kde"),
type_sim_conformalize = c("surrogate", "block-bootstrap", "bootstrap", "kde",
"fitdistr"),
ml_method = NULL,
level = 95,
B = 250L,
ml = TRUE,
stat_model = NULL,
n_clusters = 0,
clustering_dist = "euclidean",
clustering_method = "kmeans",
...
)Arguments
- y
A numeric vector or time series of class
ts- h
Forecasting horizon
- mean_model
Function to fit the mean model (default:
forecast::auto.arima)- model_residuals
Function to model the residuals (default:
forecast::thetaf)- fit_func
Fitting function for the variance model (default:
ahead::ridge)- predict_func
Prediction function for the variance model (default:
predict)- conformal
A boolean; either use conformal prediction or not
- lags_vol
Number of lags for squared residuals regression
- type_pi
Type of prediction interval ("kde", "surrogate", or "bootstrap") for volatility modeling
- type_sim_conformalize
Type of simulation for conformalization of standardized residuals ("block-bootstrap", "surrogate", "kde", "bootstrap", or "fitdistr")
- ml_method
caret package Machine learning method to use, if
fit_funcandpredict_funcaren't provided. If NULL, usesfit_funcandpredict_func. Seeunique(caret::modelLookup()$model).- level
Confidence level for prediction intervals
- B
Number of bootstrap replications or simulations
- ml
If
TRUE,fit_funcandpredict_funcare used, otherwise a statistical model instat_model- stat_model
A statistical model, e.g
forecast::thetaforforecast::auto.arima- n_clusters
Number of clusters for residuals (default is 0) for
ml == TRUEandml_methodnotNULL- clustering_dist
Only if
n_clusters > 0. If "euclidean", then mean square error, if "manhattan ", the mean absolute error is used.- clustering_method
Only if
n_clusters > 0. If "kmeans", then we have the kmeans clustering method, if "hardcl" we have the On-line Update (Hard Competitive learning) method, and if "neuralgas", we have the Neural Gas (Soft Competitive learning) method. Abbreviations of the method names are accepted.- ...
Additional parameters to be passed to
stat_model
Value
A forecast object containing predictions and prediction intervals
Examples
y <- fpp2::goog200
par(mfrow=c(2, 2))
(obj_ridge <- ahead::mlarchf(y, h=20L, B=500L, conformal=TRUE))
#> Point Forecast Lo 95 Hi 95
#> 201 534.4214 513.0126 549.3934
#> 202 536.2434 516.2732 554.8151
#> 203 536.4303 521.4627 553.5272
#> 204 536.6375 516.4953 559.5297
#> 205 536.9744 513.3215 555.6089
#> 206 536.8669 518.1739 552.4976
#> 207 539.0674 519.1329 558.1133
#> 208 537.4618 501.6435 560.6395
#> 209 540.1076 518.9179 558.1104
#> 210 536.5789 509.1344 555.3093
#> 211 540.7064 517.4373 555.1692
#> 212 543.6837 528.5274 561.0368
#> 213 543.4614 524.3126 564.4024
#> 214 544.3046 526.8850 564.2638
#> 215 544.1797 522.0492 560.9355
#> 216 544.7224 525.9826 560.9315
#> 217 545.7186 524.2705 564.2967
#> 218 546.5558 524.4958 568.5002
#> 219 546.0634 527.0138 562.9882
#> 220 548.5014 525.1078 567.3898
plot(obj_ridge)
print(mean(obj_ridge$upper - obj_ridge$lower))
#> [1] 39.69045
(obj_ridge <- ahead::mlarchf(y, h=20L, B=500L, conformal=FALSE))
#> Point Forecast Lo 95 Hi 95
#> 201 532.2466 517.5339 549.7756
#> 202 531.7224 508.7408 551.2735
#> 203 533.2419 519.0815 549.6869
#> 204 538.2220 519.6208 558.4489
#> 205 533.6841 515.4832 550.8369
#> 206 536.0713 522.1240 551.1259
#> 207 537.9695 520.3529 550.8776
#> 208 536.6699 519.0575 552.6136
#> 209 540.0722 520.8841 557.9272
#> 210 540.1827 527.2999 564.3716
#> 211 541.1687 524.6547 553.8595
#> 212 540.4668 524.1068 562.6405
#> 213 540.3615 522.8569 564.6440
#> 214 541.6808 524.9869 556.9404
#> 215 541.9843 526.4018 563.4523
#> 216 542.8743 527.5210 562.6983
#> 217 543.3401 528.7225 560.6608
#> 218 543.5208 522.6618 565.3813
#> 219 544.4165 528.6999 560.2330
#> 220 544.7700 526.8607 563.6331
plot(obj_ridge)
print(mean(obj_ridge$upper - obj_ridge$lower))
#> [1] 35.17146
(obj_ridge <- ahead::mlarchf(y, h=20L, B=500L, conformal=TRUE, stat_model=forecast::thetaf, ml = FALSE))
#> Point Forecast Lo 95 Hi 95
#> 201 538.0943 511.3731 599.5316
#> 202 532.7592 492.6637 561.5746
#> 203 535.6568 505.4333 574.0318
#> 204 536.4320 505.2108 562.3914
#> 205 538.5321 513.7271 572.9817
#> 206 538.4453 506.7213 584.0057
#> 207 538.5266 502.4655 572.0884
#> 208 540.0950 518.7978 568.7838
#> 209 539.1217 513.9889 573.2098
#> 210 540.4599 509.6873 566.7720
#> 211 542.2328 517.2629 579.0726
#> 212 542.7936 507.8171 581.9193
#> 213 542.4282 507.2424 582.2066
#> 214 546.1098 512.3549 595.3141
#> 215 544.6685 516.5238 597.2780
#> 216 541.0232 501.8039 573.0544
#> 217 540.6050 497.3601 577.0328
#> 218 545.2773 518.6021 581.7445
#> 219 546.9851 523.3497 586.1527
#> 220 547.9129 517.3082 577.3549
plot(obj_ridge)
print(mean(obj_ridge$upper - obj_ridge$lower))
#> [1] 68.34035
(obj_ridge <- ahead::mlarchf(y, h=20L, B=500L, conformal=FALSE, stat_model=forecast::thetaf, ml = FALSE))
#> Point Forecast Lo 95 Hi 95
#> 201 531.6567 514.0526 548.3243
#> 202 532.9804 511.2759 560.9922
#> 203 534.6671 518.1708 558.5576
#> 204 534.9105 511.9659 557.0808
#> 205 534.9256 518.7440 557.7363
#> 206 535.9355 518.3771 551.7302
#> 207 537.2476 520.0605 561.5851
#> 208 536.8801 518.2995 556.3820
#> 209 537.4439 520.6509 558.4829
#> 210 537.6221 522.2782 559.3128
#> 211 538.4523 522.5890 557.8461
#> 212 539.1346 523.1141 558.1896
#> 213 541.5791 518.9426 563.6007
#> 214 540.9385 521.0993 559.8572
#> 215 541.4986 521.7432 562.2131
#> 216 542.6545 527.2600 564.8567
#> 217 543.5398 525.5320 564.5464
#> 218 544.5900 527.8212 567.4387
#> 219 544.2469 523.9578 565.1556
#> 220 545.0149 525.0901 566.1182
plot(obj_ridge)
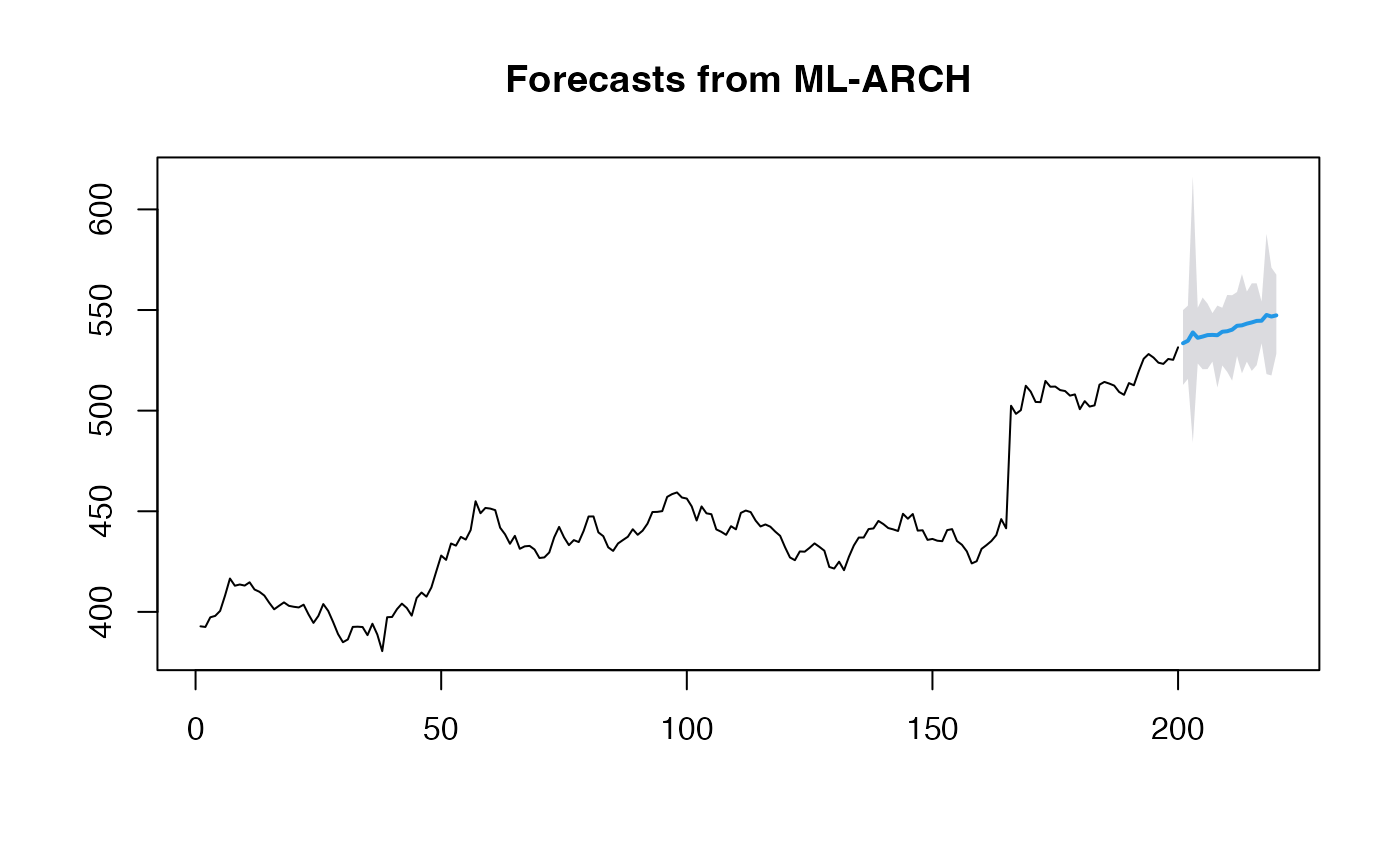 print(mean(obj_ridge$upper - obj_ridge$lower))
#> [1] 39.44909
par(mfrow=c(1, 1))
print(mean(obj_ridge$upper - obj_ridge$lower))
#> [1] 39.44909
par(mfrow=c(1, 1))